
Neuroscience has long been a field of divide and conquer. Researchers typically map specific cognitive functions to isolated brain regions—like motion to area V5 or faces to the fusiform gyrus—using models tailored to narrow experimental paradigms. While this has provided deep insights, the resulting landscape is fragmented, lacking a unified framework to explain how the human brain integrates multisensory information.
Meta’s FAIR team has introduced TRIBE v2, a tri-modal foundation model designed to bridge this gap. By aligning the latent representations of state-of-the-art AI architectures with human brain activity, TRIBE v2 predicts high-resolution fMRI responses across diverse naturalistic and experimental conditions.

The Architecture: Multi-modal Integration
TRIBE v2 does not learn to ‘see’ or ‘hear’ from scratch. Instead, it leverages the representational alignment between deep neural networks and the primate brain. The architecture consists of three frozen foundation models serving as feature extractors, a temporal transformer, and a subject-specific prediction block.
The model processes stimuli through three specialized encoders:
- Text: Contextualized embeddings are extracted from LLaMA 3.2-3B. For every word, the model prepends the preceding 1,024 words to provide temporal context, which is then mapped to a 2 Hz grid.
- Video: The model uses V-JEPA2-Giant to process 64-frame segments spanning the preceding 4 seconds for each time-bin.
- Audio: Sound is processed through Wav2Vec-BERT 2.0, with representations resampled to 2 Hz to match the stimulus frequency .
2. Temporal Aggregation
The resulting embeddings are compressed into a shared dimension and concatenated to form a multi-modal time series with a model dimension of . This sequence is fed into a Transformer encoder (8 layers, 8 attention heads) that exchanges information across a 100-second window.
3. Subject-Specific Prediction
To predict brain activity, the Transformer outputs are decimated to the 1 Hz fMRI frequency and passed through a Subject Block. This block projects the latent representations to 20,484 cortical vertices and 8,802 subcortical voxels.
Data and Scaling Laws
A significant hurdle in brain encoding is data scarcity. TRIBE v2 addresses this by utilizing ‘deep’ datasets for training—where a few subjects are recorded for many hours—and ‘wide’ datasets for evaluation.
- Training: The model was trained on 451.6 hours of fMRI data from 25 subjects across four naturalistic studies (movies, podcasts, and silent videos).
- Evaluation: It was evaluated across a broader collection totaling 1,117.7 hours from 720 subjects.
The research team observed a log-linear increase in encoding accuracy as the training data volume increased, with no evidence of a plateau. This suggests that as neuroimaging repositories expand, the predictive power of models like TRIBE v2 will continue to scale.
Results: Beating the Baselines
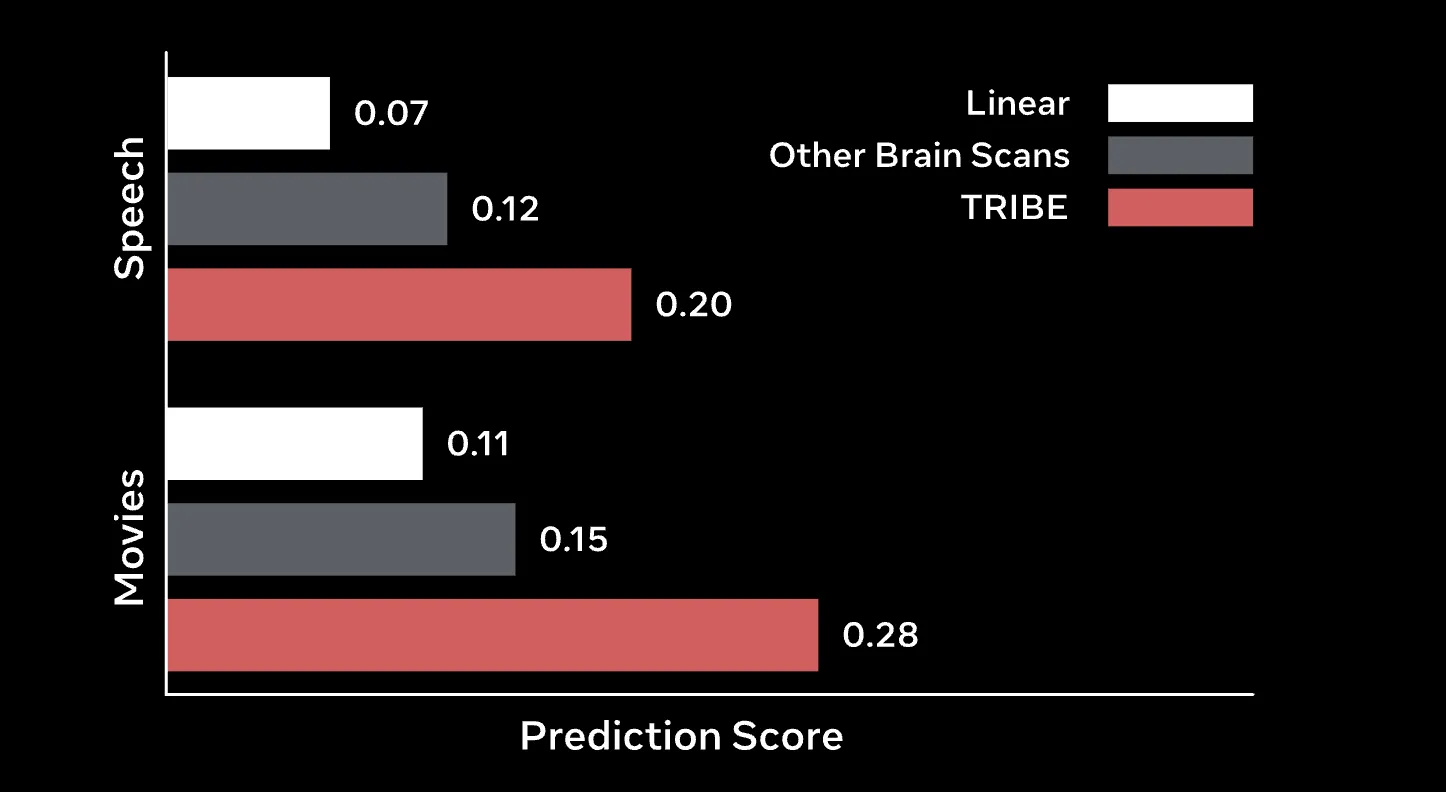
TRIBE v2 significantly outperforms traditional Finite Impulse Response (FIR) models, the long-standing gold standard for voxel-wise encoding.
Zero-Shot and Group Performance
One of the model’s most striking capabilities is zero-shot generalization to new subjects. Using an ‘unseen subject’ layer, TRIBE v2 can predict the group-averaged response of a new cohort more accurately than the actual recording of many individual subjects within that cohort. In the high-resolution Human Connectome Project (HCP) 7T dataset, TRIBE v2 achieved a group correlation near 0.4, a two-fold improvement over the median subject’s group-predictivity.
Fine-Tuning
When given a small amount of data (at most one hour) for a new participant, fine-tuning TRIBE v2 for just one epoch leads to a two- to four-fold improvement over linear models trained from scratch.
In-Silico Experimentation
The research team argue that TRIBE v2 could be useful for piloting or pre-screening neuroimaging studies. By running virtual experiments on the Individual Brain Charting (IBC) dataset, the model recovered classic functional landmarks:
- Vision: It accurately localized the fusiform face area (FFA) and parahippocampal place area (PPA).
- Language: It successfully recovered the temporo-parietal junction (TPJ) for emotional processing and Broca’s area for syntax.
Furthermore, applying Independent Component Analysis (ICA) to the model’s final layer revealed that TRIBE v2 naturally learns five well-known functional networks: primary auditory, language, motion, default mode, and visual.

Key Takeaway
- A Powerhouse Tri-modal Architecture: TRIBE v2 is a foundation model that integrates video, audio, and language by leveraging state-of-the-art encoders like LLaMA 3.2 for text, V-JEPA2 for video, and Wav2Vec-BERT for audio.
- Log-Linear Scaling Laws: Much like the Large Language Models we use every day, TRIBE v2 follows a log-linear scaling law; its ability to accurately predict brain activity increases steadily as it is fed more fMRI data, with no performance plateau currently in sight.
- Superior Zero-Shot Generalization: The model can predict the brain responses of unseen subjects in new experimental conditions without any additional training. Remarkably, its zero-shot predictions are often more accurate at estimating group-averaged brain responses than the recordings of individual human subjects themselves.
- The Dawn of In-Silico Neuroscience: TRIBE v2 enables ‘in-silico’ experimentation, allowing researchers to run virtual neuroscientific tests on a computer. It successfully replicated decades of empirical research by identifying specialized areas like the fusiform face area (FFA) and Broca’s area purely through digital simulation.
- Emergent Biological Interpretability: Even though it’s a deep learning ‘black box,’ the model’s internal representations naturally organized themselves into five well-known functional networks: primary auditory, language, motion, default mode, and visual.
Check out the Code, Weights and Demo. Also, feel free to follow us on Twitter and don’t forget to join our 120k+ ML SubReddit and Subscribe to our Newsletter. Wait! are you on telegram? now you can join us on telegram as well.








